In the current age of the COVID-19 pandemic, we find ourselves inundated with headline articles and millions of daily social media posts on the novel coronavirus. For the frontline healthcare worker, the massive amount of online information can be overwhelming, and for the general public, it often feels as though they are being plunged into anxiety and panic. The goal of these articles is to help the end-user understand some fundamental topics as they relate to pandemics, viruses, and how we as a society deal with them. Our hope is that these posts will make it easier to navigate through the current spread of medical information.
COVID-19, Part I – Virus, Virology, and Vaccines
Coronaviruses and COVID-19
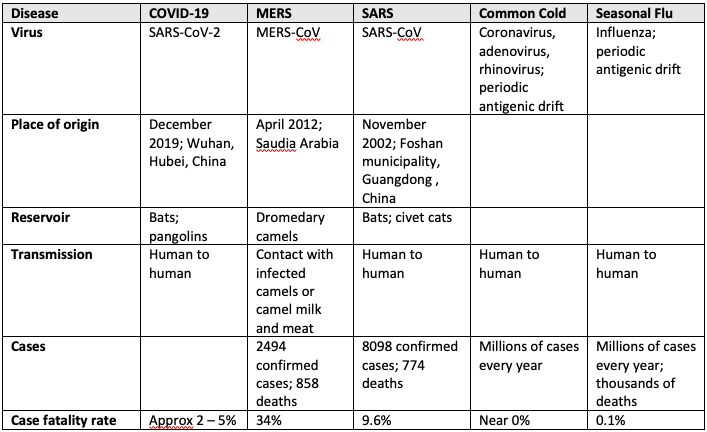
Coronaviruses (CoV) are non-segmented positive sense RNA viruses whose name derives from their characteristic crown-like appearance. They can be divided into four subgroups: α, β, γ, and δ. The α– and β-CoV have the ability to infect mammals, while the γ- and δ-CoV have known to infect birds. Previously, 6 human coronaviruses (2 α-CoV and 4 β-CoV) have been discovered. These include two α–CoV (229E, NL63) and two β-CoV (HKU1, OC43), strains that are responsible for about 20% of the “common cold” (the other 80% are due to various strains of rhinoviruses and adenoviruses). Discovered in 2002 and 2012 respectively, the other two β-CoVs, SARS-CoV and MERS-CoV, have caused severe respiratory infections in humans resulting in worldwide epidemics. The following table shows some similarities between commonly known viruses.
The newly discovered SARS-CoV2 virus is 79.5% genetically identical to the SARS-CoV virus and 96% identical to a known bat CoV (RaTG13). Most recently, researchers in Guangzhou, China, have found 99% genetic similarity to a CoV found in pangolins, an ant-eating mammal often used in traditional Chinese medicine. Based on this knowledge, it has been presumed that the SARS-CoV2 might be transmitted from bats via unknown intermediate hosts such as the pangolins to infect humans.
Understanding the origins and transmission of a virus is important as it has major public health implications. Because of the previous SARS outbreaks and the current pandemic, the Chinese government has temporarily banned the trade of wild animals and currently drafting a permanent law to prevent the trade of certain wild animals in wet markets. These wet markets are often linked to the origin of viruses where many reservoir animals (civet cats for the SARS-CoV, pangolins for the SARS-CoV-2) are kept in close proximity and unsanitary conditions. The Arabian Peninsula has seen strict regulation of camel movement and trade after MERS-CoV hit the region. While many argue these public health interventions are necessary, these policies are often met with friction as they challenge the very nature of century-old cultures of public trade and food consumption in the Asian and Middle Eastern countries.
Relevance to selected targeted therapies
Because the genetic material is similar to SARS-CoV, and because it widely researched in the early 2000s, we know that the novel virus has RNA that encodes for 4 similar structural proteins: spike (S), membrane (M), nucleocapsid (N), envelop (E) proteins. Notably the S protein binds to host cell receptor ACEII, a critical step for virus entry. The binding of ACEII has clinical implications as it is abundantly present on alveolar epithelial cells (specifically type 2 pneumocytes) and enterocytes of small intestines, explaining the respiratory and GI symptoms that may accompany the SARS-CoV-2 infection. There is also emerging evidence that once the virus enters alveolar cells, a series of immune and cytokine responses are triggered that may ultimately contribute to sustained lung injury in critically ill patients.
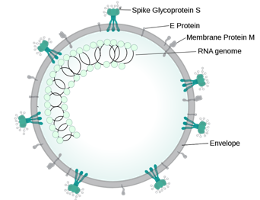
Wikipedia.org
How does our knowledge of the coronavirus relate to the recently proposed therapies? As mentioned above, the SARS-CoV-2 virus has a single RNA genome that can directly produce its proteins once it has entered the host cell. It uses the host cell’s ribosomes to translate the genetic material into proteins. These proteins assemble together to form virus particles that are eventually transported out of the host cell to infect other cells. This self-replication machinery is common to many viruses and explains why they are so effective at causing an overwhelming response.
An important step in the viral life cycle is its reliance on an enzyme called protease. There are a few proteases in the SARS-CoV-2 genome, such as PLpro and 3CLpro, that cleave proteins to make them “mature” and ready to use. A common antiviral currently being studied is Kaletra, or Lopinavir/Ritonavir, which is a protease inhibitor used to target HIV proteases. Despite evidence that it has in vitro activity against SARS-CoV and MERS-CoV, clinical efficacy has not been definitely established.
Another critical step in its life cycle is the use of an RNA-dependent RNA polymerase, or RdRp, a highly conserved protein throughout viruses that is used to replicate its own genome. This is necessary to continue the genetic footprint of viruses and continue the self-replication machinery. Another common antiviral, Remdesivir, an adenosine nucleotide analog, interferes with the action of RdRP. This virus was initially developed as a treatment for Ebola and Marburg, however recent in vitro studies have demonstrated possible action against a variety of viruses including SARS-CoV-2. Unfortunately, despite a recent compassionate use cohort study, the data presented offered limited information regarding its efficacy and safety profile for the treatment of COVID-19.
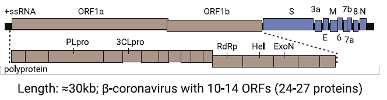
The coronavirus genome: notable genes include proteases (PLpro, 3CLpro), the RNA-dependent RNA polymerase (RdRp), and antigenic proteins (S or spike, M or membrane, N or nucleocapsid, E or envelope) – wikipedia.org
COVID-19 vaccine development
The WHO declared a global pandemic on March 11, 2020 and many countries had begun imposing strict control measures to limit social contact and some going onto unprecedented national lockdowns. However, even with the most effective and at times draconian containment strategies, the absence of herd immunity against COVID-19 means that the virus could easily resurge when interventions are relaxed. Current prediction models suggest this may very well be the case if countries loosen their control measures too early. This is particularly why novel viruses can often overwhelm medical systems, especially if they have high infectivity and transmissibility. While there are multiple proposed treatment options against the virus as above, many have failed to demonstrate a mortality benefit.
Fortunately, since the genetic sequence of SARS-CoV-2 was published on January 11, 2020, many companies are racing to develop a vaccine against the virus. To appreciate the scale of the COVID-19 vaccine landscape, it is important to understand how vaccine technologies are developed and the success of the existing influenza vaccines.
Influenza vaccines
Modern flu vaccines were first developed in the 1930s and protected US troops against the flu during World War II. Nowadays, a new vaccine is developed each year to combat the latest flu strains. There exist more than 100 national influenza centres around the world that conduct year-round influenza surveillance. They are involved with testing thousands of virus samples from patients. The labs send representative viruses (ie. those that have similar antigenic representation in multiple patients in their region) to five WHO Collaborating Centres for Reference and Research on Influenza, located in: Atlanta, London, Melbourne, Tokyo, and Beijing. The WHO then reviews the results of surveillance, lab, and clinical studies and provide recommendations on the composition of the vaccine. They meet biannually to participate in the seasonal vaccine virus selection for the Southern and Northern Hemispheres. Generally speaking, vaccines are developed using three different technologies:
- Egg-based vaccines, the most common and oldest way of manufacturing vaccines, uses inactivated or live attenuated viruses that are replicated in fertilized hen’s eggs. The candidate viruses are injected into the eggs and incubated for several days before the fluid is harvested and virus antigen is purified.
- Cell-based vaccines are made by using inactivated viruses that have been grown in cultured mammalian cells.
- Recombinant vaccines, the newest technology, are created using viral genetic material (ie. DNA or RNA) targeted for specific proteins on the virus. For-example the hemagglutinin (HA) protein found in influenza is commonly used as a target. The genetic material can either be injected directly into humans, or used to replicate the protein in host cell lines that is then injected into humans. Either method provides antigens (ie. HA) for the body to mount an immune response against.
SARS-COV-2 vaccine pipeline
So what about SARS-CoV-2? The focus of vaccine production has shifted away from the traditional egg-based methods as this technology takes the longest. This was a hard lesson that many countries learned when it led to delays in distributing a vaccine for the H1N1 pandemic in 2009. By the time a vaccine was mass produced, H1N1 was largely “under control”. Additionally, with strained healthcare systems, rising cases and fatality rates, national lockdowns and blunted economies, the pressure is on for countries to develop a vaccine as soon as possible. Thus, scientists are turning towards recombinant technology that, once developed, can be upscaled rapidly.
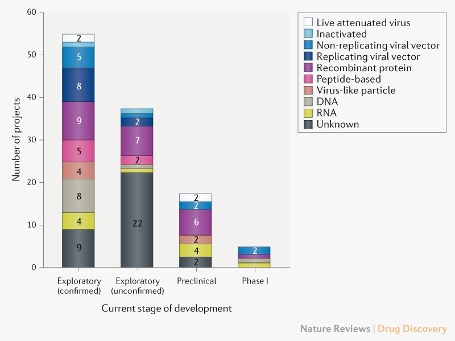
Pipeline of COVID-19 vaccine candidates by technology platform. From Nature Reviews.
As of April 2020, there are 115 vaccine candidates for SARS-CoV-2. While a few include traditional methods such as using live attenuated and inactivated viruses, a majority of these candidates include genetic material (DNA or mRNA), virus particles, protein segments, viral vectors, and recombinant proteins. The idea is to present either the genetic material or virus particles to immune cells that can develop antibodies against antigens specific to the SARS-CoV-2 virus. A main target for many of these vaccines is the spike protein, the viral antigen that is used for host cell entry.
One of the silver linings from this pandemic is the fact that vaccine research and development is being mobilized at a rapid pace. To illustrate one of the hopeful stories, scientists at the NIH and Moderna, a Massachusetts-based biotech company, used the genetic instructions for making the spike protein into an instruction molecule, or messenger RNA. Their mRNA-1273 candidate is currently in its Phase I trial and is designed to instruct cells to mass-produce the spike protein, which then presents themselves to other immune cells in hopes of producing antibodies and effectively protecting one against SARS-CoV-2. If scientists can demonstrate these novel platforms are safe and effective, they can use the same structural backbone and manipulate genetic instructions to create other viral proteins against the human immune system. There is evidence to suggest that the current SARS-CoV-2 virus, like other coronaviruses and influenzae viruses, may develop mutations and re-infect in the future. If we can develop robust recombinant vaccine technology, it won’t be too long until these candidates can provide a promising therapy against such viruses that may show antigenic drift.
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382(8):727–733. Yin Y, Wunderink RG, MERS, SARS and other coronaviruses as causes of pneumonia. Respirology. 2018 Feb; 23(2):130-137. Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL Nature. 2020 Mar; 579(7798):270-273. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 395 (10223): 497–506. Leung K, Wu JT, Liu D, Leung GM. First-wave COVID-19 transmissibility and severity in China outside Hubei after control measures, and second-wave scenario planning: a modelling impact assessment. Lancet. April 2020. Guo RY, Cao QD, Hong ZS, Tan YY, Chen SD, Jin HJ, Tan KS, Wang DY, Yan Y. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak – an update on the status. Mil Med Res. 2020; 7:11. Chu DK, Poon LL, Gomaa MM, et al. MERS coronaviruses in dromedary camels, Egypt. Emerg Infect Dis. 2014;20(6):1049–1053. Thanh Le T, Andreadakis Z, Kumar A, et al. The COVID-19 vaccine development landscape [published online ahead of print, 2020 Apr 9]. Nat Rev Drug Discov. 2020. Grein J, Ohmagari N, Shin D, et al. Compassionate use of remdesivir for patients with severe Covid-19. N Engl J Med. 2020. Epub. DOI: 10.1056/NEJMoa2007016 Chu CM, Cheng VC, Hung IF et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004; 59:252-6. (PubMed 14985565) (DOI 10.1136/ thorax.2003.012658) Cao B, Wang Y, Wen D et al. A Trial of Lopinavir-Ritonavir in Adults Hospitalized with Severe Covid-19. N Engl J Med. 2020; (PubMed 32187464) (DOI 10.1056/NEJMoa2001282) https://www.cnn.com/2020/04/10/asia/china-wildlife-law-coronavirus-intl-hnk/index.html https://www.cdc.gov/flu/prevent/how-fluvaccine-made.htm https://www.who.int/blueprint/priority-diseases/key-action/novel-coronavirus-landscape-ncov.pdfReferences


Nicely done and not overwhelming.